NEW DIAGNOSTIC TECHNOLOGIES

APTAMERS, WHAT ARE THEY?
Aptamers are short segments of nucleic acids (oligonucleotides) or peptidic molecules capable of binding to a target molecule with high specificity and affinity due to their three-dimensional structure.
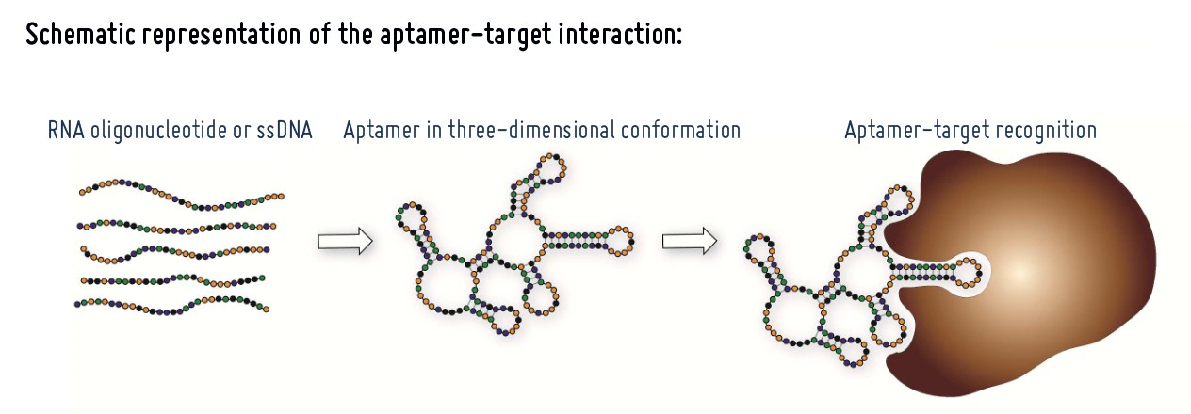
Adapted from Darmostuk M et al.(2015)
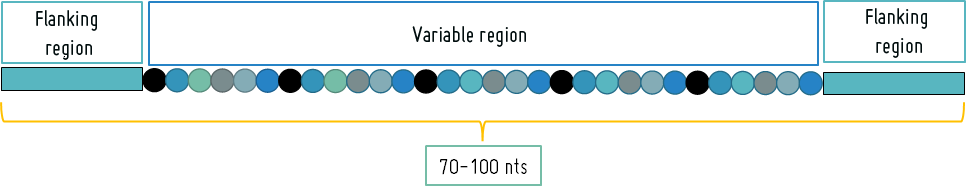
They have a length of between 70 and 100 nucleotides, with a central region of variable size (between 30 and 60 nucleotides) of random sequence and two flanking regions of known sequence for their amplification by PCR. They can adopt globular structures that allow them to recognize molecules by binding to them in a stable and specific manner.
APPLICATIONS OF APTAMERS
Aptamers have a wide range of applications, both in diagnosis and treatment, serving as an alternative to the use of monoclonal antibodies. Additionally, they offer advantages over antibodies: they are easier to produce, can be easily modified, can recognize any target (immunogenic and non-immunogenic) rather than just immunogenic ones like antibodies do, and do not elicit an immune response in the body.
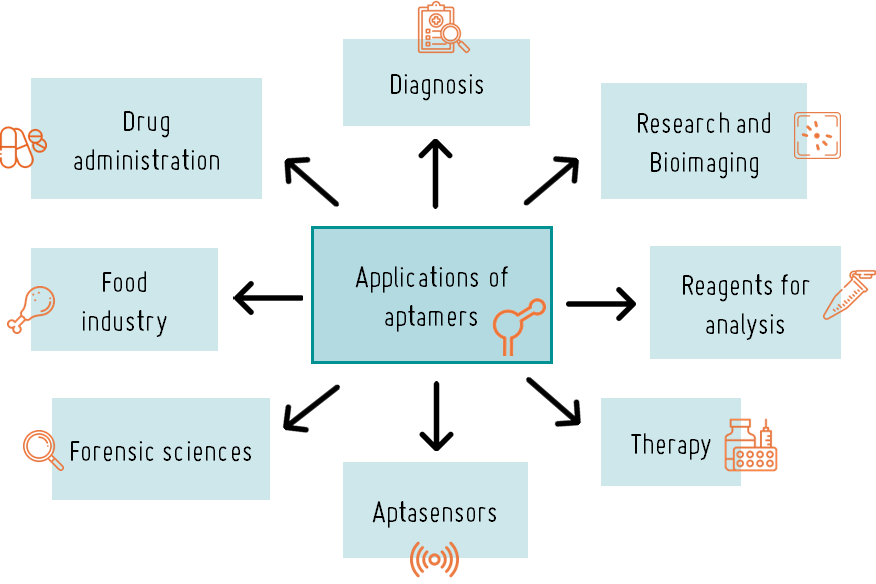
However, while there are several ongoing clinical trials for their diagnostic and therapeutic applications, the use of aptamers in the clinical setting is still emerging, indicating that there is still much to explore in this area.
CHARACTERIZATION OF APTAMERS
Aptamers are selected through the SELEX method (Systematic Evolution of Ligands by EXponential enrichment). This method involves exposing an initial library of oligonucleotides to our target molecule. Through this process, some oligonucleotides, due to their three-dimensional structure, will bind to the target while the rest are washed away. The bound oligonucleotides are then recovered and amplified via PCR to generate a library of oligonucleotides with higher affinity for the target molecule in each selection round. In the final selection round, the oligonucleotides are sequenced to identify candidate aptamers with high affinity for our target molecule.
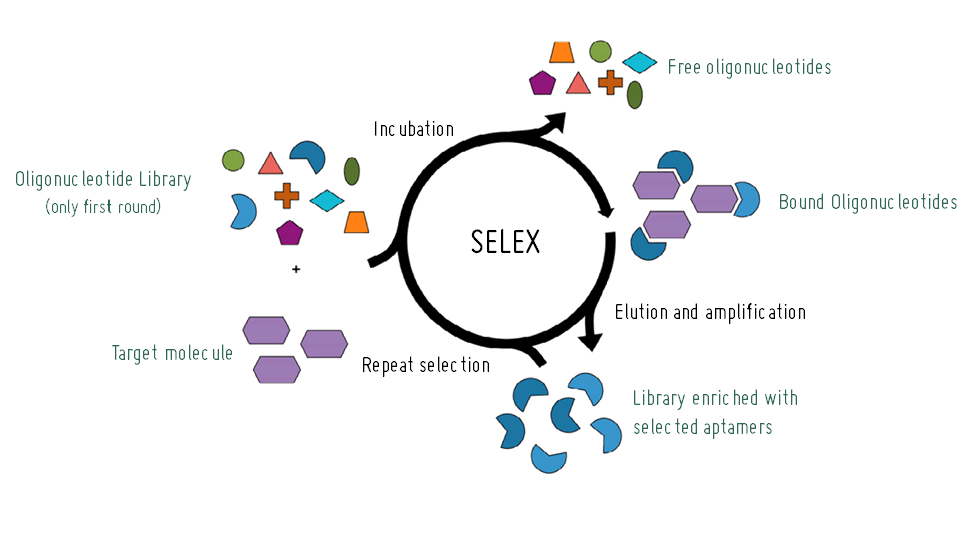
Adapted from Zhang, et al. (2019)
ELONA ASSAYS WITH APTAMERS
An ELONA is a bioassay in which aptamers labeled with enzymes are bound to a target of interest. It allows determining whether the aptamers selected for molecule recognition actually bind to the target and with what sensitivity, affinity, and specificity, in order to select the best candidates for use in the diagnosis or treatment of a specific condition (for example: tumor marker recognition or HIV diagnosis in our case).
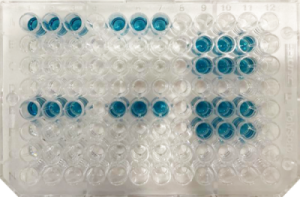

The ELONA is conducted in a plate with 96 wells. In a direct ELONA, the target molecule is placed at the bottom of the well (in this example, the HIV reverse transcriptase protein). Subsequently, another protein called BSA (bovine serum albumin) is added to cover any gaps that may have remained at the bottom of the well. Then, the aptamer labeled with digoxigenin is added, which is a small molecule that binds to the anti-digoxigenin antibody attached to the HRP enzyme (horseradish peroxidase). When ABTS (HRP substrate) is added, if the aptamer has bound to the target molecule, a reaction occurs that produces a color change in the well's liquid, and its intensity or absorbance can be measured. If, however, the aptamer is not specific and does not bind to the target molecule, there will be no reaction and no color will be generated.
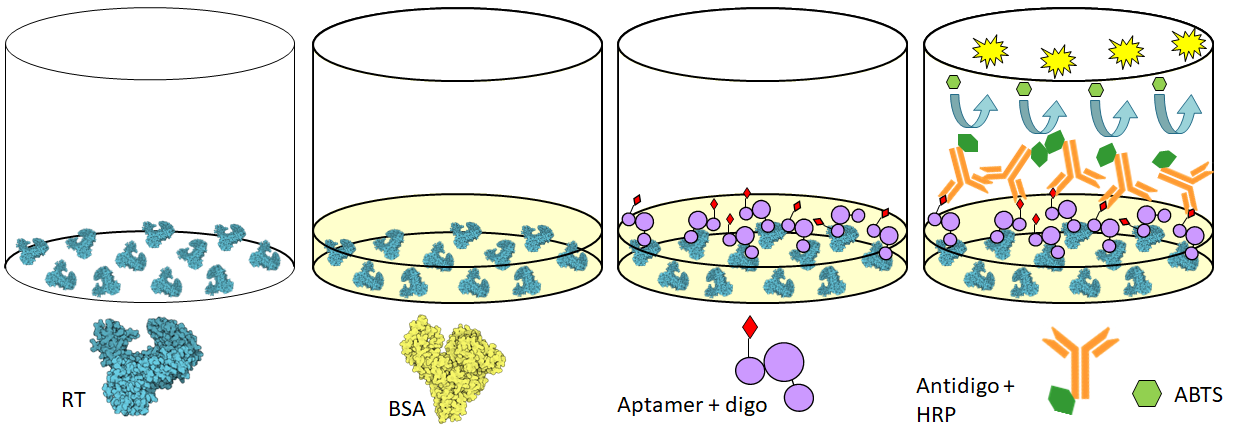
There are also other types of ELONA called sandwich ELONA in which a combination of aptamers (one capturing aptamer and one detecting aptamer) is used to detect the target molecule, increasing the sensitivity of the experiment.
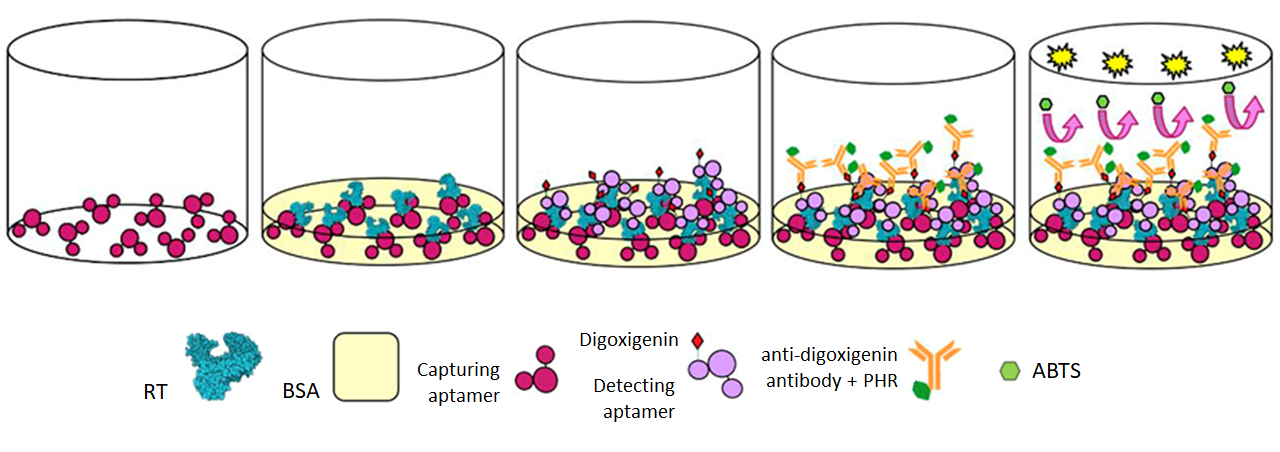
APTAMERS IN HIV
Current HIV diagnostic techniques only allow virus detection after the first 10 days following infection using molecular techniques (which directly recognize the virus through its RNA or proteins) and from 3 weeks using serological techniques (which recognize specific antibodies against the virus) or 2 weeks if using state-of-the-art ELISAs (4th and 5th generation) that detect antibodies and p24 protein of the virus. 40% of HIV transmissions occur during the acute phase of the infection in the absence of treatment due to the high viral load in the first 3 weeks after infection, before antibodies against the virus begin to be generated. Therefore, it is necessary to develop new molecular diagnostic techniques capable of recognizing the virus from the onset of infection. Aptamers are an interesting option for the development of these new techniques, as they are able to recognize viral proteins that are more abundant than the genetic material of the virus.
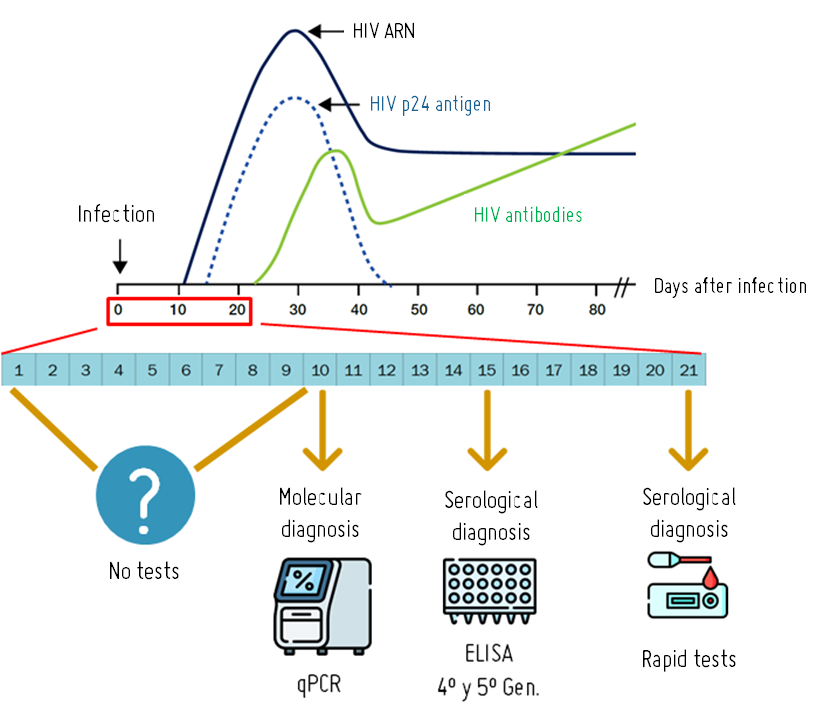
ELONAs present certain limitations in detection sensitivity, often requiring a high viral load for accurate results. Therefore, there is a need for the development of new, highly sensitive detection methods.
PROJECTS IN PROGRESS WITH APTAMERS
In EPIMOLVIH, since 2018, we have been working with aptamers. We have identified peptides from 5 highly conserved viral proteins across all HIV variants to select different aptamers for the molecular detection of HIV, evaluating their sensitivity, affinity, and specificity for future diagnostic implementation. Concurrently, we are testing new detection technologies based on aptamers to enable early diagnosis within the first 10 days of infection.

You can find more information about our projects with aptamers here.
Do you want to collaborate with us to continue advancing in the research of new technologies for HIV diagnosis?
You too can contribute to advancing research in aptamers and new HIV diagnostic technologies that allow for the detection of infection in the early days, enabling early treatment initiation and helping to prevent new infections. Would you like to collaborate? Join our Teaming group for just one euro per month or donate whatever amount you consider appropriate as an individual or company. Together for advancing research in new HIV diagnostic technologies!



